Development of a generalisable tryptophan-optimised quenchbody biosensor based on a synthetic nanobody library
Abstract
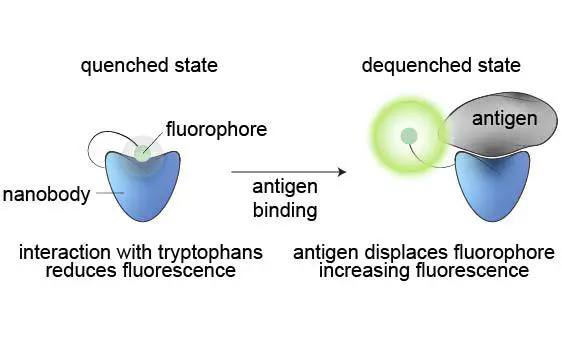
Quenchbodies are antibodies labelled with a fluorophore that increases in intensity upon antigen binding, which makes them promising candidates for the development of diagnostic or other immunoassays requiring antigen quantification. Quenchbodies based on camelid nanobodies are particularly attractive for rapid development of immunoassays against molecular antigens of interest, due to their small size, ease of expression, high stability, rapid evolvability, and amenability to protein engineering. However, current nanobody-based quenchbodies typically display only modest fluorescence increases when binding to protein antigens (<= 1.1{\textendash}1.7-fold), with few examples reaching > 2-fold. In contrast to single-chain variable region (scFv; 32 kDa) or antigen-binding fragment (Fab; 50 kDa) quenchbodies, where semi-conserved tryptophans at the VH-VL interface quench the fluorophore, here we show that tryptophan residues in the nanobody complementarity-determining regions (CDRs) are critical to fluorescence antigen detection. This informed the development of a synthetic convex-binding nanobody library containing conserved tryptophans in the CDRs, which was used for the in vitro selection and affinity maturation of novel quenchbodies against human inflammatory cytokine interleukin 6 (IL6). Initial in silico modelling and biochemical analyses showed that existing nanobodies for maltose-binding protein (MBP) and lysozyme could be converted into quenchbodies (Qb-MBP and Qb-Lys), with fluorescence fold-increases of 1.5 and 1.3, respectively, when recognising their cognate antigen. Rational mutational substitution of tryptophans into the CDR-region of the quenchbodies resulted in fluorescence fold-increases of 1.9 and 1.6 for Qb-MBP (Y59W/Y114W) and Qb-Lys (Y110W), respectively, supporting the importance of CDR-based tryptophans in the nanobody quenchbody mechanism. A synthetic nanobody library containing conserved CDR-based tryptophans was then created, selected, and affinity matured using a completely in vitro SNAP-display system, yielding quenchbodies against interleukin-6 (IL6) with enhanced fluorescence fold-increases of 1.5{\textendash}2.4 (EC50 = 20{\textendash}1,113 nM binding affinity). The ease and speed by which nanobody-based quenchbodies can be discovered using this completely in vitro selection strategy based on a single synthetic library, makes this a very attractive approach to develop immunoassays for detection of a wide range of molecular targets, including proteins.